---
title: "Begun, the Package Manager Wars, Have"
author: "Tinashe M. Tapera"
date: "2026-04-07"
image: aotcql.jpg
categories: [r, package-management, rv, uvr, rust]
title-block-banner: true
---
There’s a quiet war brewing in the data science community, and no, it’s not about which AI model hallucinates the least.
It’s about package management.
As data science matures, reproducibility and portability are no longer optional.
Models, analyses, and pipelines need to run reliably across laptops, clusters, and
cloud environments. Tools like VSCode devcontainers, GitHub Codespaces, Google Colab,
and Posit Cloud have made development more portable—but they’ve also made dependency
management more critical than ever.
At the same time, the field is still figuring out its standards. What used to be a
niche concern (package conflicts, environment drift, etc.) is now something even
undergraduate students are taught to think about. And with that has come an explosion
of tools: `pip`, `virtualenv`, `renv`, `conda`, `mamba`, `docker`, `nix`, `uv`, `pak`,
and more. Each brings a different philosophy about how environments should be defined,
resolved, and reproduced.
The abundance of tools is powerful, but it’s also overwhelming. It’s increasingly
common for data scientists to stitch together multiple tools, often without a clear
model of how they interact.
So how should you navigate this landscape?
A good starting point is scope. A small script may only need `pip` or `renv`, while a
production pipeline might justify Docker or Nix. Personally, I believe the key is to
choose something that scales slightly beyond your predicted needs, so you don’t incur
technical debt as your work grows.
Next is control. Are you working locally, on a shared HPC cluster, or in the cloud? Do
you have root access? Different tools assume different levels of control, and
mismatches here are a common source of frustration.
Finally, consider usability and community support. The most powerful tool is not
always the most obviously practical, especially when collaborating. I recommend
favouring tools that your end-users and collaborators can understand, adopt, and debug
collectively without too much headache.
I’m not going to claim there’s a single “best” solution. But I do think there’s a
useful way to think about these tools, and it leads naturally to the newest entrant
worth paying attention to: rv.
## `rv` is Pretty Fast

Because `rv` was built in Rust, it is designed to be fast and efficient. If you want to understand why,
just watch this video of how memory works in the Rust programming language: [https://www.youtube.com/watch?v=5C_HPTJg5ek](https://www.youtube.com/watch?v=5C_HPTJg5ek).
While some packages can take up to and even beyond 60 seconds with
`install.packages()`, installing stuff in `rv` often takes a fraction of
that time. This is mainly because, being a Rust-based software, `rv` is
able to manage internal data structures and operations at lightning speed!
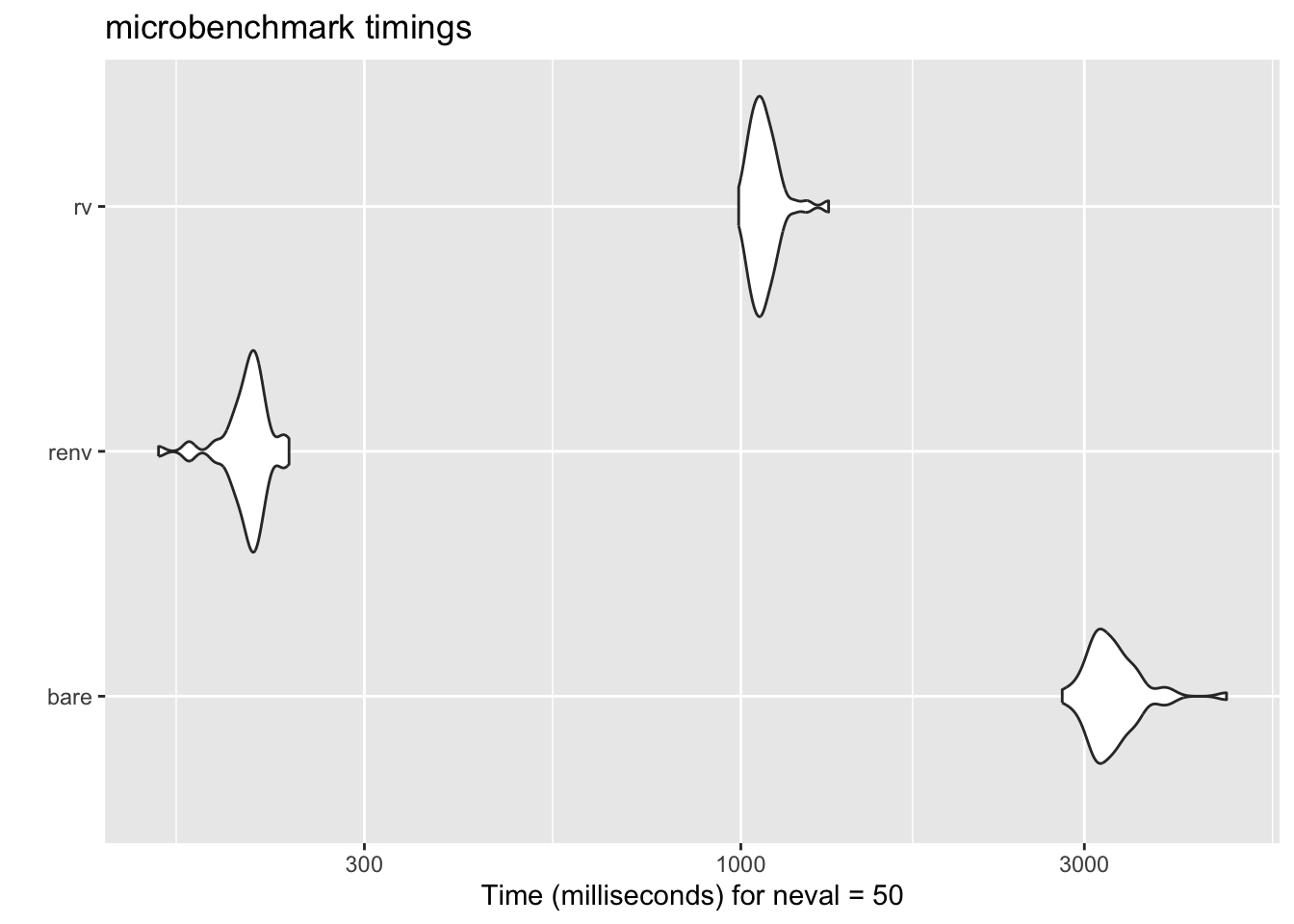
To demonstrate this, I set up a microbenchmark to compare the time it takes
to install a few popular packages using `install.packages()`, `renv`, and `rv`.
Even with the necessary `init` and `configure` steps, the results were quite
impressive.
Thanks ChatGPT for setting up a microbenchmark:
```{r, message=FALSE, warning=FALSE}
library(microbenchmark)
library(tidyverse)
# microbenchmark_install.R
# -----------------------------
# Configuration
# -----------------------------
pkgs <- c("tidyverse", "tidymodels", "data.table")
repos <- "https://cloud.r-project.org"
# helper to create clean temp dir
make_clean_dir <- function(prefix) {
path <- file.path(tempdir(), paste0(prefix, "_", as.integer(Sys.time())))
dir.create(path, recursive = TRUE, showWarnings = FALSE)
normalizePath(path)
}
# helper to run R in isolated process
run_r_script <- function(code, lib, wd) {
script <- tempfile(fileext = ".R")
writeLines(code, script)
system2(
command = file.path(R.home("bin"), "Rscript"),
args = c(script),
env = c(
paste0("R_LIBS_USER=", lib),
paste0("R_LIBS=", lib)
),
stdout = TRUE,
stderr = TRUE,
wait = TRUE
)
}
# -----------------------------
# 1. Bare R
# -----------------------------
bench_bare <- function() {
wd <- make_clean_dir("bare")
lib <- file.path(wd, "library")
dir.create(lib, recursive = TRUE)
code <- sprintf('
dir.create("%s", recursive = TRUE, showWarnings = FALSE)
.libPaths("%s")
install.packages(c(%s), repos = "%s", Ncpus = parallel::detectCores())
',
lib,
lib,
paste(sprintf('"%s"', pkgs), collapse = ", "),
repos
)
message("Running bare R install...")
t <- system.time(run_r_script(code, lib, wd))
return(t)
}
# -----------------------------
# 2. renv
# -----------------------------
bench_renv <- function() {
wd <- make_clean_dir("renv")
code <- sprintf('
cd %s
install.packages("renv", repos = "%s")
renv::init(bare = TRUE)
install.packages(c(%s), repos = "%s", Ncpus = parallel::detectCores())
',
wd,
repos,
paste(sprintf('"%s"', pkgs), collapse = ", "),
repos
)
message("Running renv install...")
t <- system.time({
system2(
file.path(R.home("bin"), "Rscript"),
args = c("-e", shQuote(code)),
stdout = TRUE,
stderr = TRUE
)
})
# delete the renv folder to clean up
unlink(file.path(wd, "renv"), recursive = TRUE, force = TRUE)
return(t)
}
# -----------------------------
# 3. rv (CLI)
# -----------------------------
bench_rv <- function() {
wd <- make_clean_dir("rv")
message("Running rv install...")
t <- system.time({
old <- setwd(wd)
on.exit(setwd(old), add = TRUE)
system("rv init", intern = TRUE)
# explicitly configure a repository
system(
paste(
"rv configure repository add --url https://packagemanager.posit.co/cran/latest cran-latest"
),
intern = TRUE
)
system(
sprintf("rv add %s", paste(pkgs, collapse = " ")),
intern = TRUE
)
system("rv sync", intern = TRUE)
# delete the rv folder to clean up
unlink(file.path(wd, "rv"), recursive = TRUE, force = TRUE)
})
t
}
# -----------------------------
# Run benchmarks
# -----------------------------
results <- microbenchmark(
bare = bench_bare(),
renv = bench_renv(),
rv = bench_rv(),
times = 50
)
autoplot(results)
```
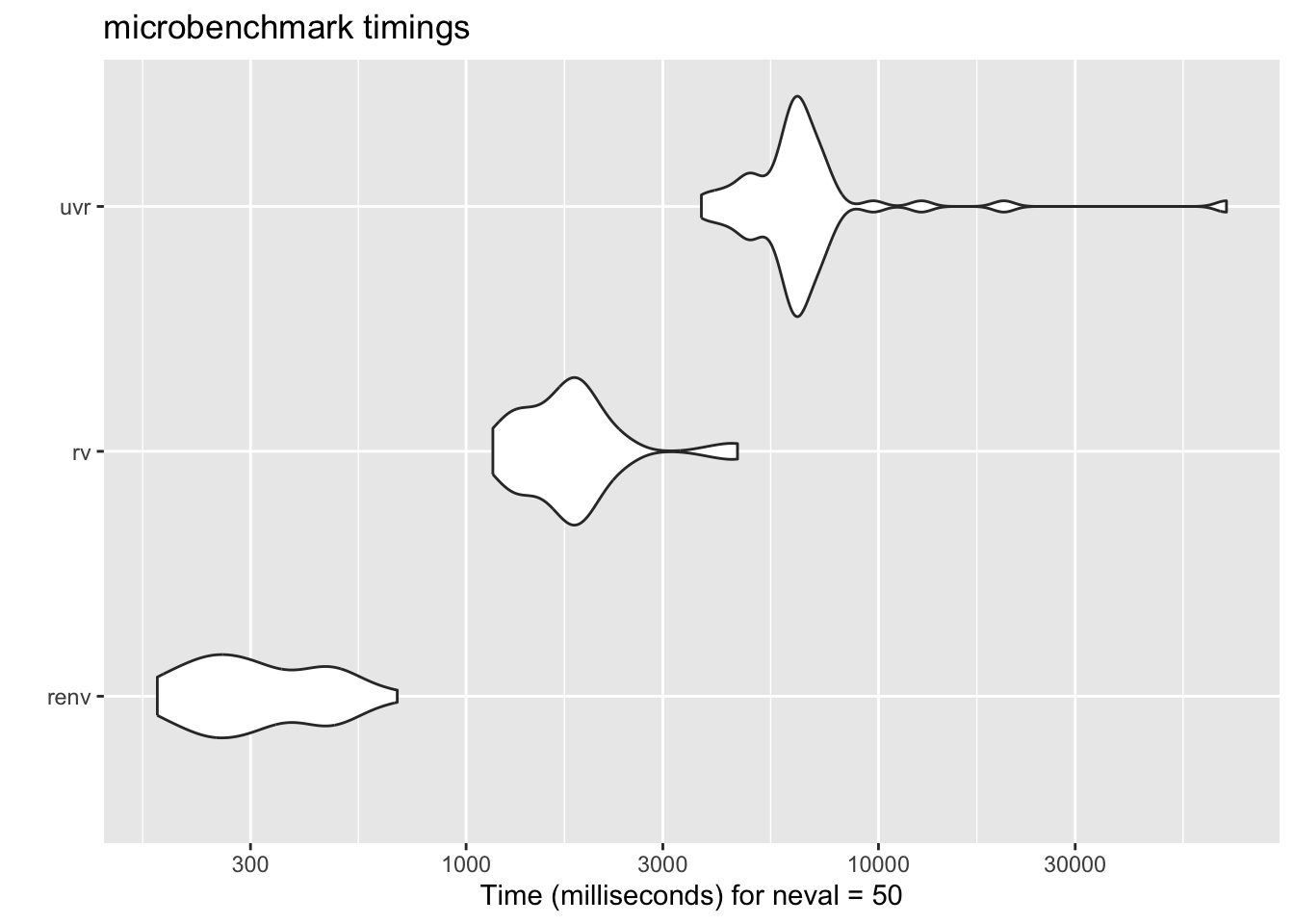
::: {.callout-important}
This isn't a perfect comparison, of course, because base R doesn't
do the environment resolution that `renv` and `rv` do, but it still gives a
good sense of the relative performance. To compare, let's add another
contender: `uvr` by [`nbafrank`](https://github.com/nbafrank/uvr), which is another Rust-based package manager. I
hadn't learned about `uvr` at the time of writing, but it seems to be a
similar tool with a similar design philosophy. So the benchmark should
apply to both `rv` and `uvr`, and it will be interesting to see how they
compare in terms of speed and usability.
```{r}
bench_uvr <- function() {
wd <- make_clean_dir("uvr")
message("Running uvr install...")
t <- system.time({
old <- setwd(wd)
on.exit(setwd(old), add = TRUE)
system("uvr init", intern = TRUE)
system(
sprintf("uvr add %s", paste(pkgs, collapse = " ")),
intern = TRUE
)
system("uvr sync", intern = TRUE)
# delete the uvr folder to clean up
unlink(file.path(wd, "uvr"), recursive = TRUE, force = TRUE)
})
t
}
```
```{r, message=FALSE, warning=FALSE}
results <- microbenchmark(
#bare = bench_bare(),
renv = bench_renv(),
rv = bench_rv(),
uvr = bench_uvr(),
times = 50
)
autoplot(results)
```
:::
## `rv` is Encapsulated

Something that I would not have argued in my early R days is that it's
a good thing that `rv` (and to its credit, `renv` as well) is encapsulated
and technically separate from the interface you use to interact with R. For
a noob, this can seem daunting — why would you want to EXIT R to do something
related to your R project? But the more comfortable I get with the terminal,
the more I appreciate the separation of concerns. This framework allows you
to manage your packages and environments as a separate task, allowing you to
focus and narrow down on specific problems. You don't have to worry if `conda`,
your `conda r-base`, or some other factor is interfering with your R session.
Instead, you install `rv` once, and use it only when you begin a project
and install a package. When you advance through your analysis, you simply
leave R, use `rv` to add or remove packages, and then return to R to continue your
work. This forces you to evaluate your package management decisions deliberately,
and should reveal any borked environments _before_ you get into the weeds of your
analysis. Better still, using tools like Quarto is more likely to work on the first
try since the `rv` binary is just sitting in your regular path and the library
itself is in your project folder.
## `rv` is Cross-Platform

Again, due to its Rust-based architecture, `rv` is designed to be cross-platform and
work seamlessly on different operating systems, including Windows, macOS, and Linux.
This means that you can use `rv` to manage your packages and environments regardless
of the platform you're working on, which can be especially useful if you're
collaborating with others who may be using different operating systems. With `rv`, you
can ensure that your project is reproducible and consistent across different
environments, making it easier to share your work and collaborate with others in the
data science community.
## `rv` is Declarative

_What does it mean for a program to be declarative?_ _Why does it even matter?_
Consider your typical workflow with base R and no additional tools. First,
you start R, then you load your data. You find that you want to plot something,
so you install `ggplot2` with `install.packages()`. Then you load `ggplot2` with
`library()`, and make your plot. You save your script, but you haven't recorded
what packages are used or how they are installed. _No worry,_ you may think.
That's what `renv` is for! You can just run `renv::snapshot()` and it will record the
state of your packages in the project, and then you can share that with your
collaborators. Problem solved?
Not so fast.
The issue with building up your package library step by step is that by the time
you have installed package number 14, written 200 lines of code, fit 4 models, and
made 6 plots, you have _iteratively_ changed the state of your software environment in
a way that is not guaranteed to be reproducible. Sure, you recorded what packages
and versions you have at the end, but what if installing package number 4 actually
caused a problem with package number 2 by updating an upstream dependency?
This is why _declarative_ package management is so powerful. With `rv`, you are forced
to declare the final build state of your project, and anytime `rv` is called,
it attempts to rebuild that state from scratch, with exact package versions and
dependencies respected.
## Conclusion

I'm loving the idea of a declarative, Rust-based package manager for R, and I think
`rv` has a lot of potential to be a game-changer in the data science community.
Rust in general offers significant performance improvements, and when applied to
package management, it may provide a more efficient and reliable solution.
I'm going to keep an eye out for any obvious pitfalls, of course, but for now,
I'm excited to see how `rv` evolves and how it can help data scientists manage their packages and environments more effectively.
::: {.callout-tip}
Bonus 1: If you are developing packages, the `rv` team has recommended that
you use `rv` like [this](https://github.com/A2-ai/scicalc/blob/main/rproject.toml)
to install the package in your local library.
Bonus 2: If you want to use `rv` in a Quarto project, you have to let
Quarto know not to render anything in the `rv` library. Do this
by adding the following to your YAML front matter:
```yaml
project:
type: website # or whatever kind of project you have
render:
- "*.qmd"
- "!rv/"
```
:::